RNA World — Codon Bias & Amino Acid Symmetry
✦ Context
This project was carried out as a research collaboration project with the University of Aberdeen (School of Natural & Computing Sciences), under the supervision of Dr. Juliano Morimoto, as part of the Origins of Life project investigation. The work explores how nucleotide composition constraints shape amino acid distributions in synthetic RNA sequences — a question directly relevant to early molecular evolution.
✦ Tools & Skills
✦ Key Findings
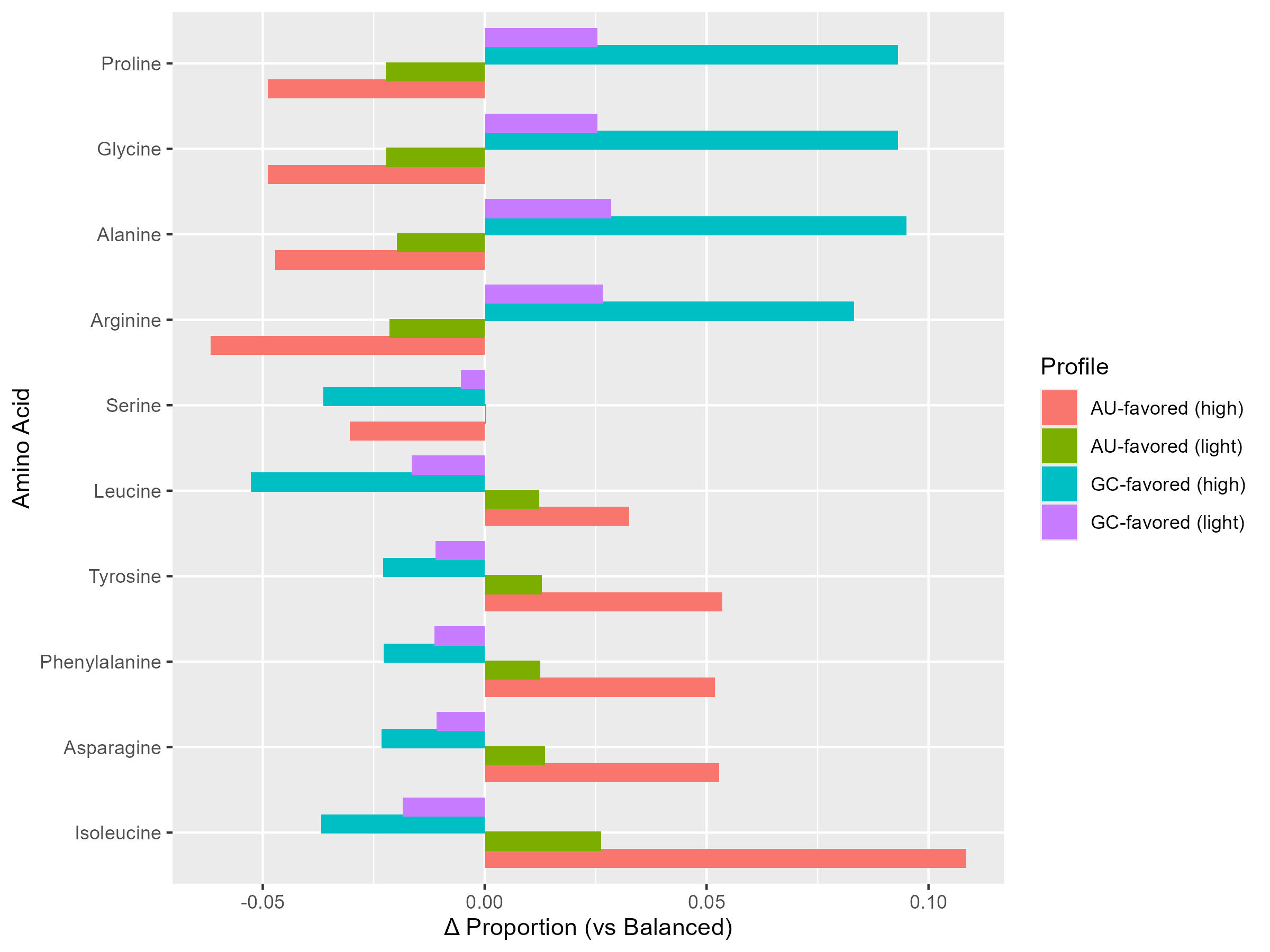
Strong positive correlation between codon redundancy and amino acid frequency (ρ up to 0.75), particularly under balanced and GC-rich conditions. Nucleotide bias shifts amino acid composition predictably — GC-rich profiles favour Proline, Glycine and Alanine, while AU-rich conditions enrich Isoleucine and Asparagine. Despite these biases, strand compositional symmetry is preserved as sequence length increases.
✦ Report
